Слайд 2

Site-specific recombination
Moves specialized nucleotide sequence (mobile genetic elements) between non-homologous sites
within a genome.
Transpositional site-specific recombination
Conservative site-specific recombinatinon
Слайд 3

Transpositional site-specific recombination
Modest target site selectivity and insert mobile genetic elements
into many sites
Transposase enzyme cuts out mobile genetic elements and insert them into specific sites.
Слайд 4
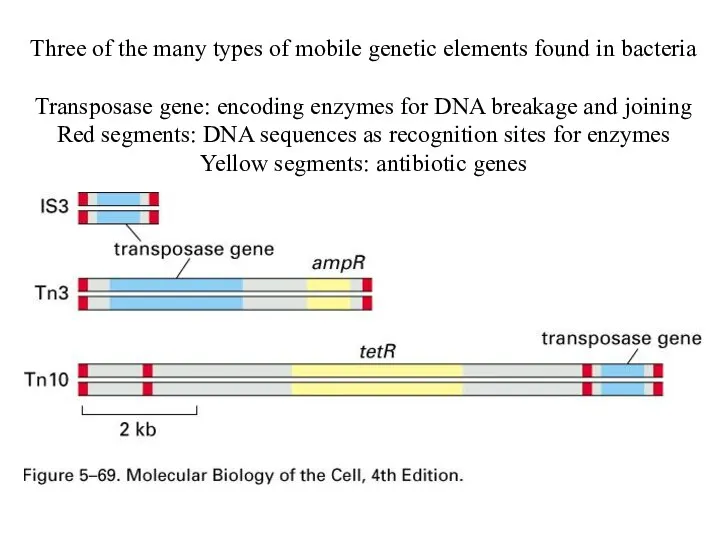
Three of the many types of mobile genetic elements found in
bacteria
Transposase gene: encoding enzymes for DNA breakage and joining
Red segments: DNA sequences as recognition sites for enzymes
Yellow segments: antibiotic genes
Слайд 5

Слайд 6
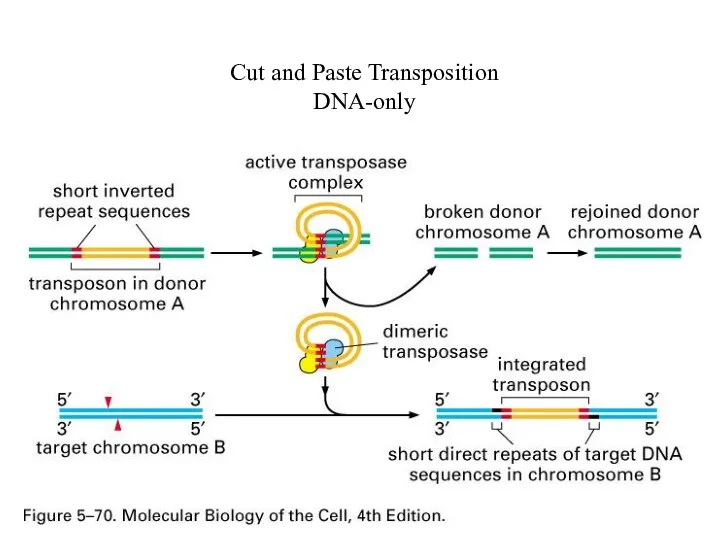
Cut and Paste Transposition
DNA-only
Слайд 7
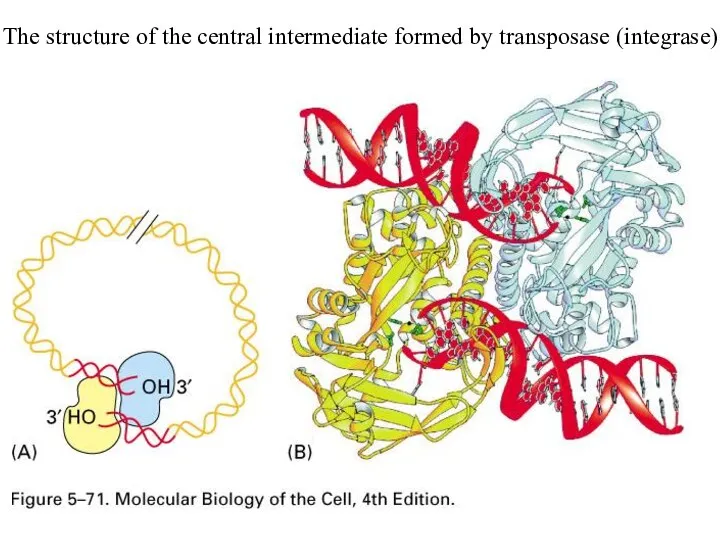
The structure of the central intermediate formed by transposase (integrase)
Слайд 8
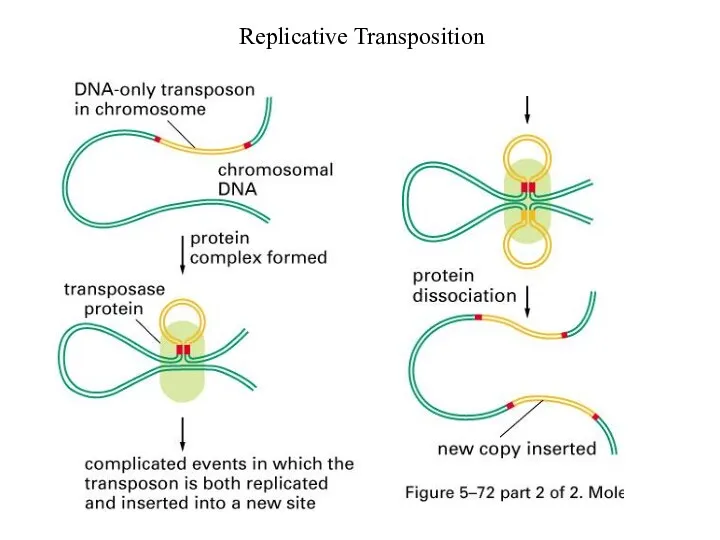
Replicative Transposition
Слайд 9
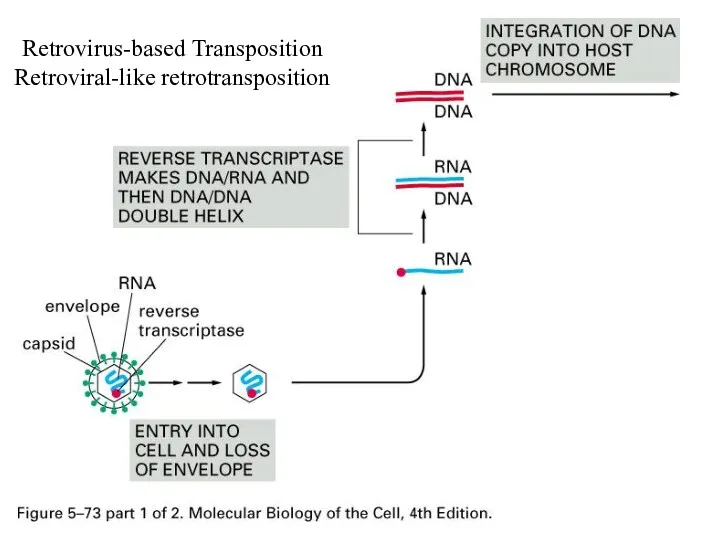
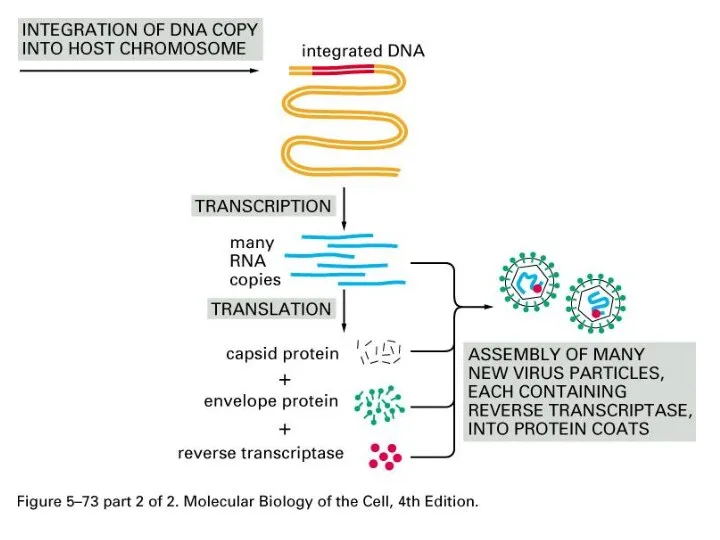
Retrovirus-based Transposition
Retroviral-like retrotransposition
Слайд 10
Слайд 11
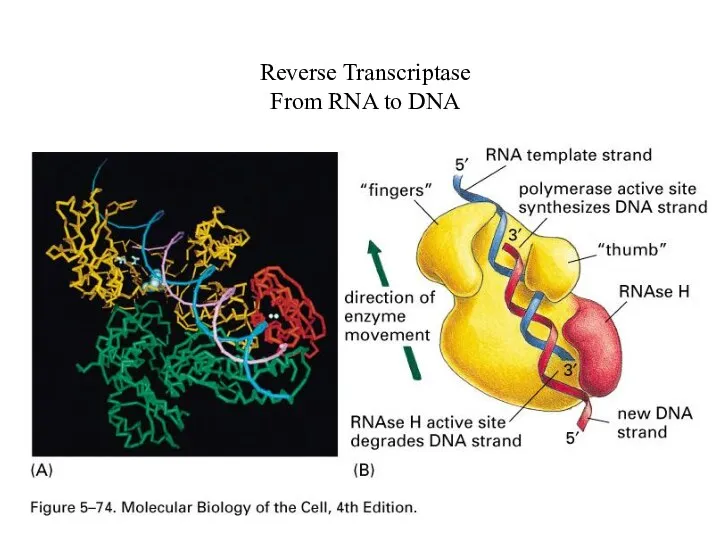
Reverse Transcriptase
From RNA to DNA
Слайд 12
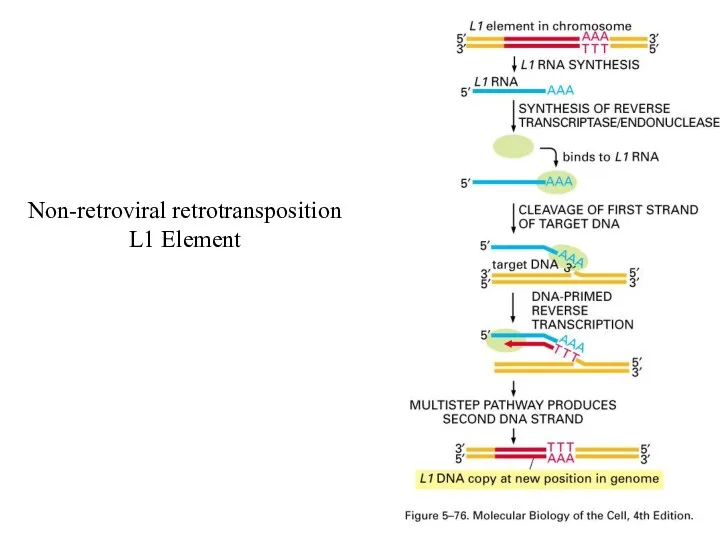
Non-retroviral retrotransposition
L1 Element
Слайд 13
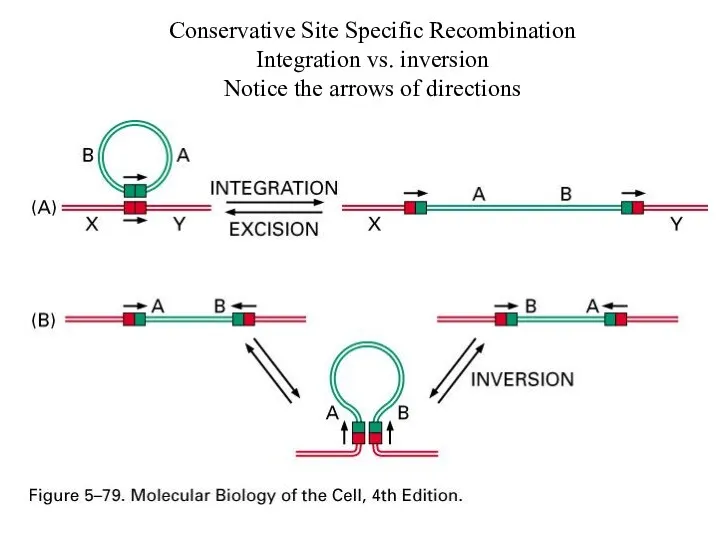
Conservative Site Specific Recombination
Integration vs. inversion
Notice the arrows of directions
Слайд 14
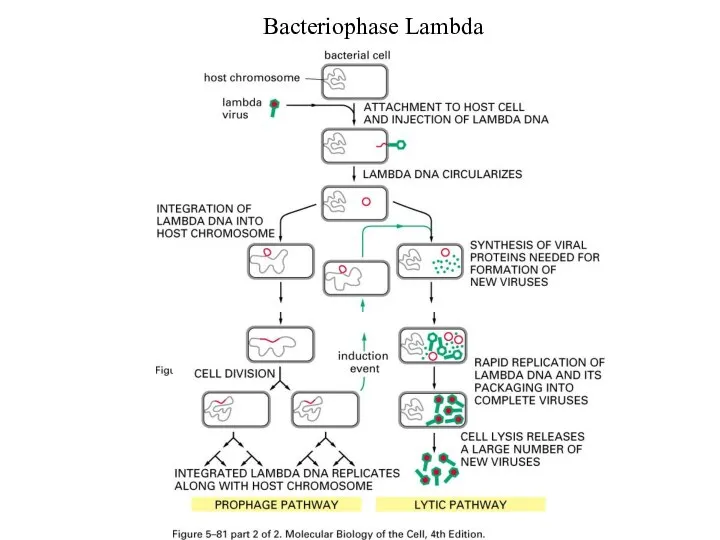
Слайд 15
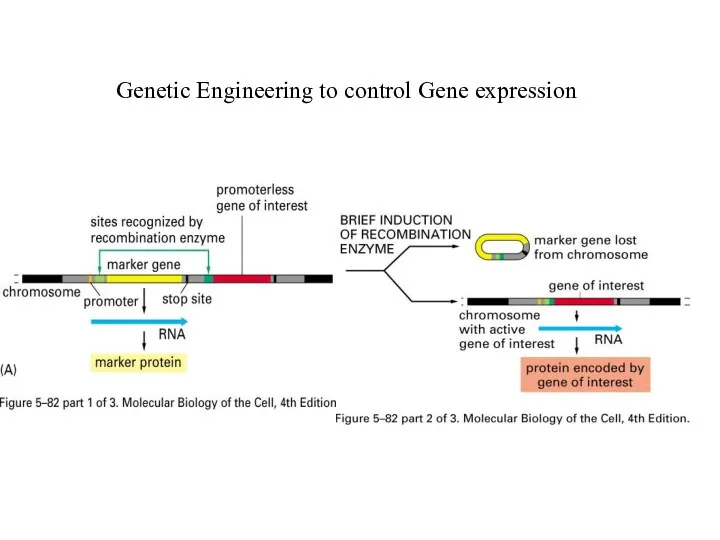
Genetic Engineering to control Gene expression
Слайд 16

Summary
DNA site-specific recombination
transpositional; conservative
Transposons: mobile genetic elements
Transpositional: DNA only transposons, retroviral-like
retrotransposons, nonretroviral retrotransposons
Слайд 17

How Cells Read the Genome: From DNA to Protein
1. Transcription
2. RNA
Modification and Splicing
3. RNA transportation
4. Translation
5. Protein Modification and Folding
Слайд 18
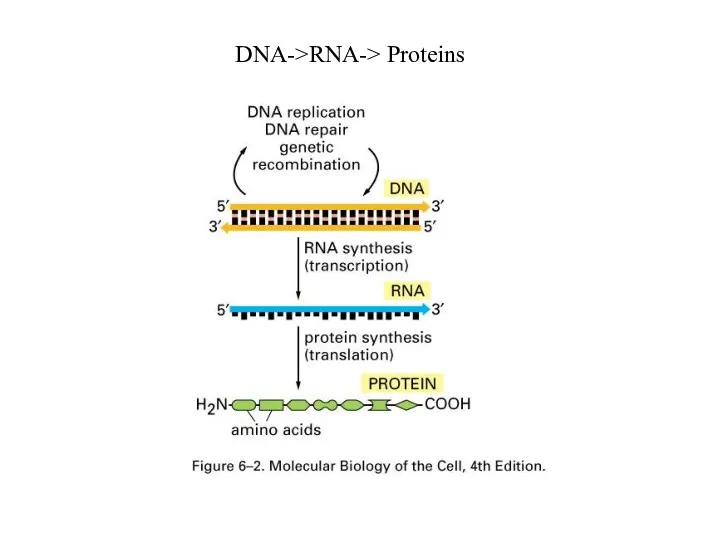
Слайд 19
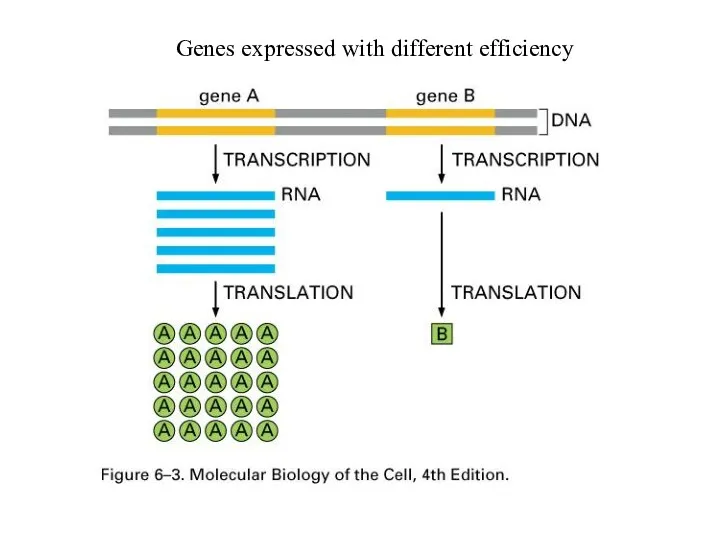
Genes expressed with different efficiency
Слайд 20
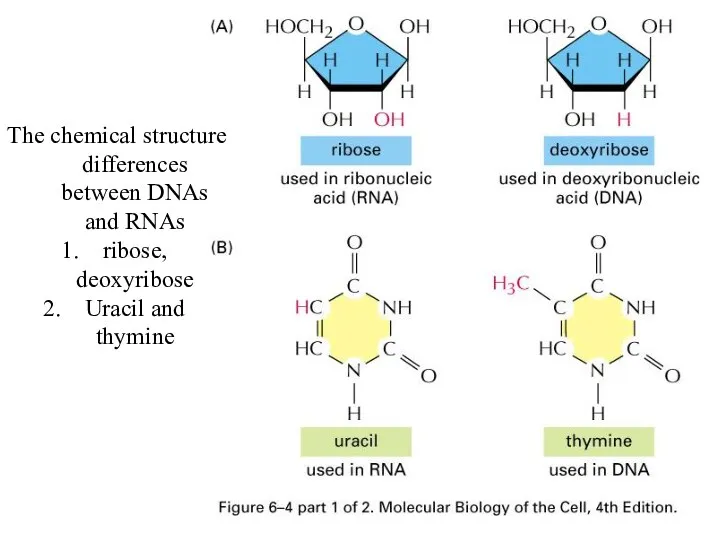
The chemical structure differences between DNAs and RNAs
ribose, deoxyribose
Uracil and thymine
Слайд 21
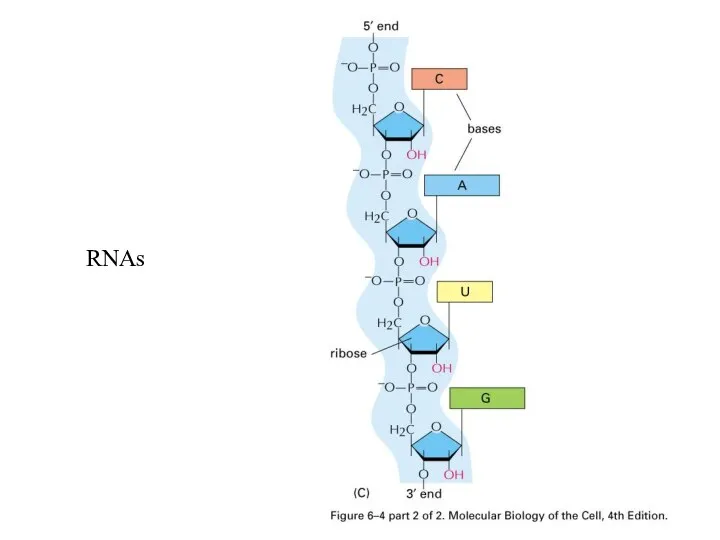
Слайд 22
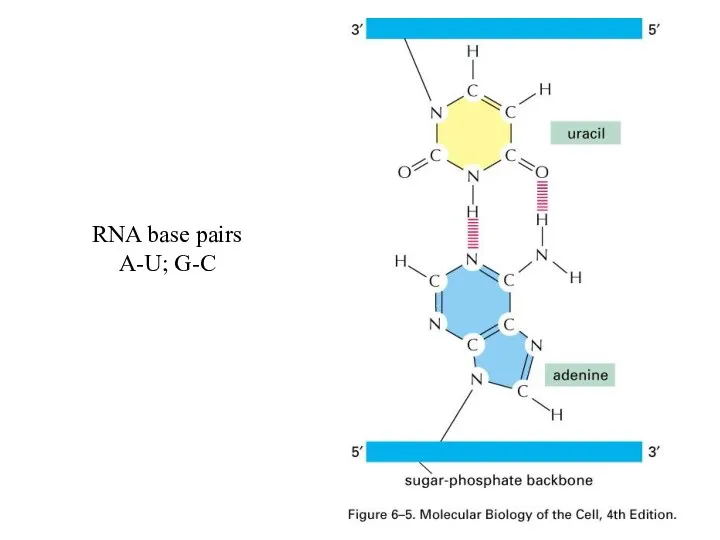
Слайд 23
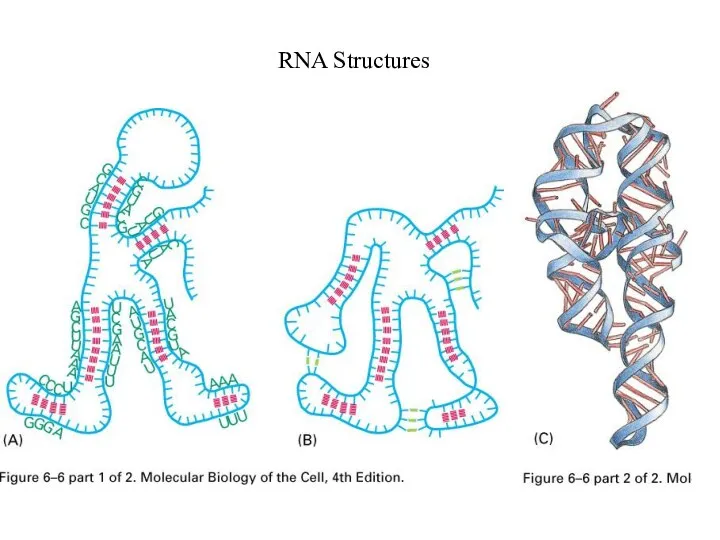
Слайд 24
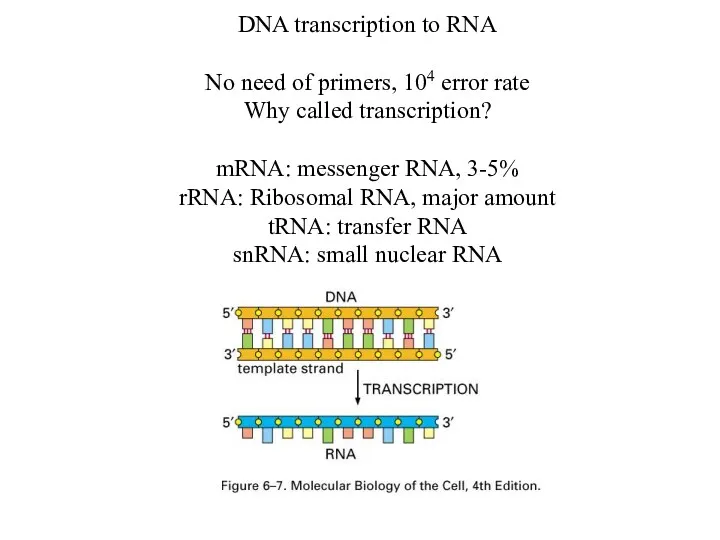
DNA transcription to RNA
No need of primers, 104 error rate
Why called
transcription?
mRNA: messenger RNA, 3-5%
rRNA: Ribosomal RNA, major amount
tRNA: transfer RNA
snRNA: small nuclear RNA
Слайд 25
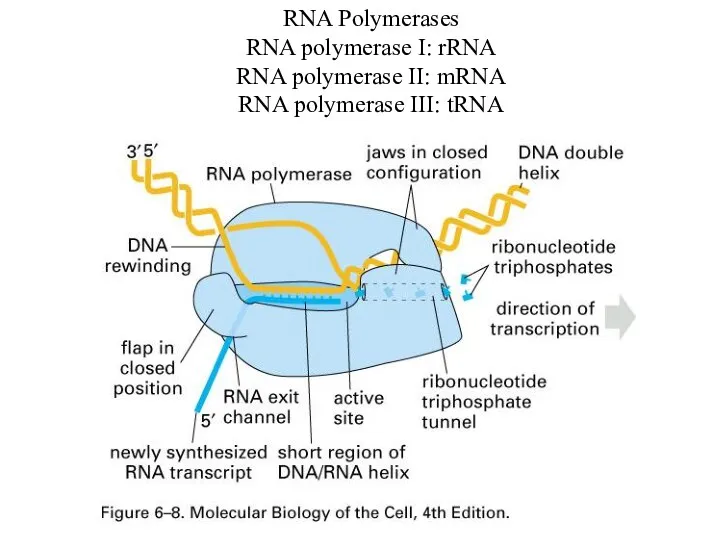
RNA Polymerases
RNA polymerase I: rRNA
RNA polymerase II: mRNA
RNA polymerase III: tRNA
Слайд 26
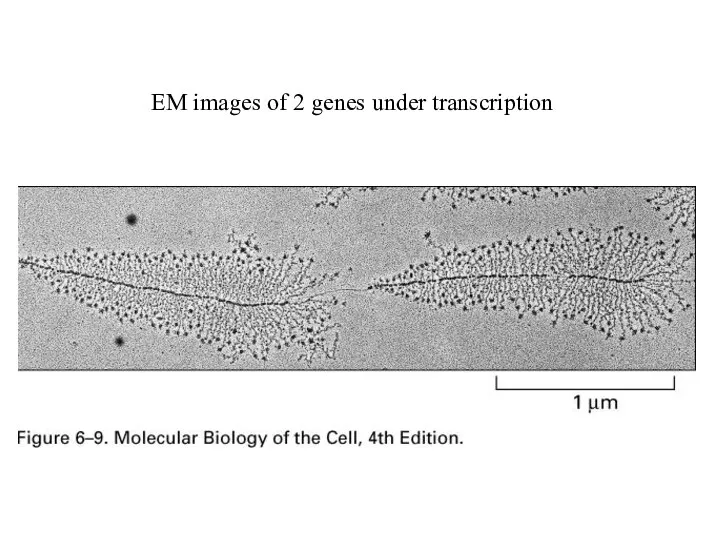
EM images of 2 genes under transcription
Слайд 27
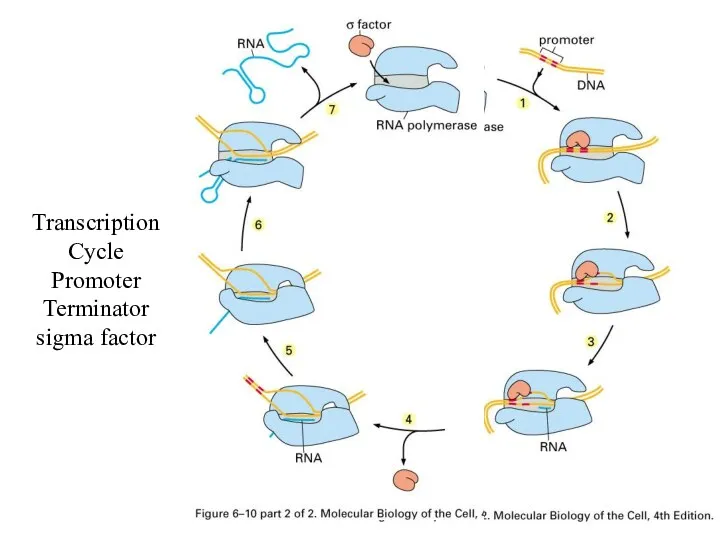
Transcription Cycle
Promoter
Terminator
sigma factor
Слайд 28
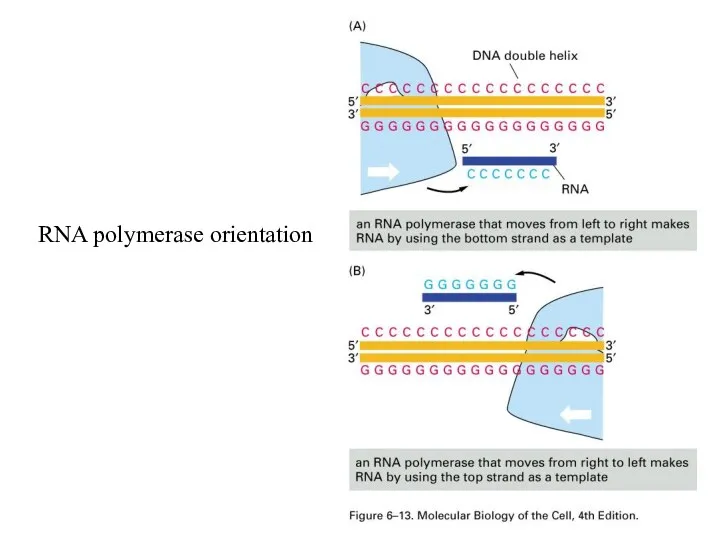
RNA polymerase orientation
Слайд 29
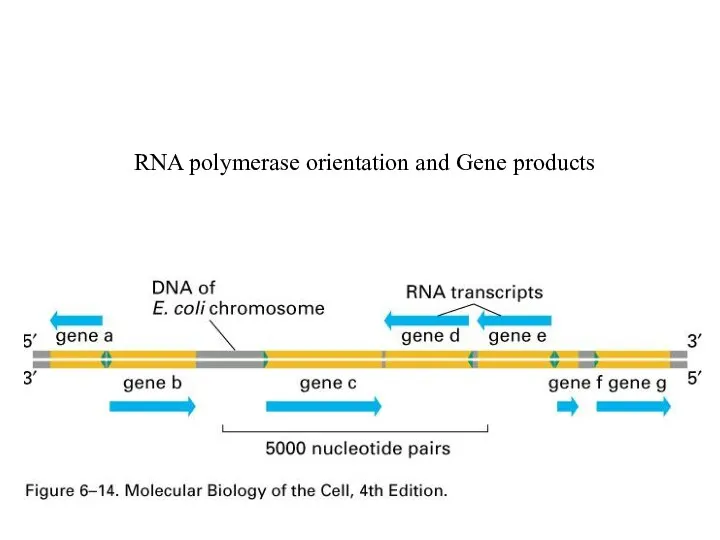
RNA polymerase orientation and Gene products
Слайд 30
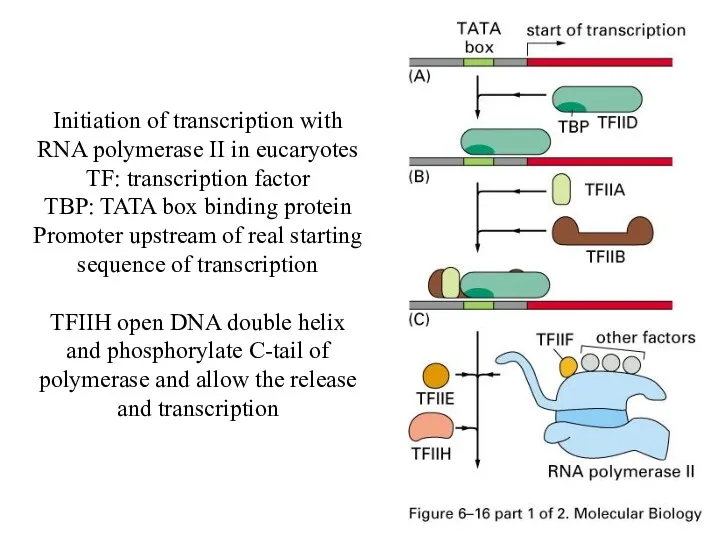
Initiation of transcription with RNA polymerase II in eucaryotes
TF: transcription factor
TBP:
TATA box binding protein
Promoter upstream of real starting sequence of transcription
TFIIH open DNA double helix and phosphorylate C-tail of polymerase and allow the release and transcription
Слайд 31
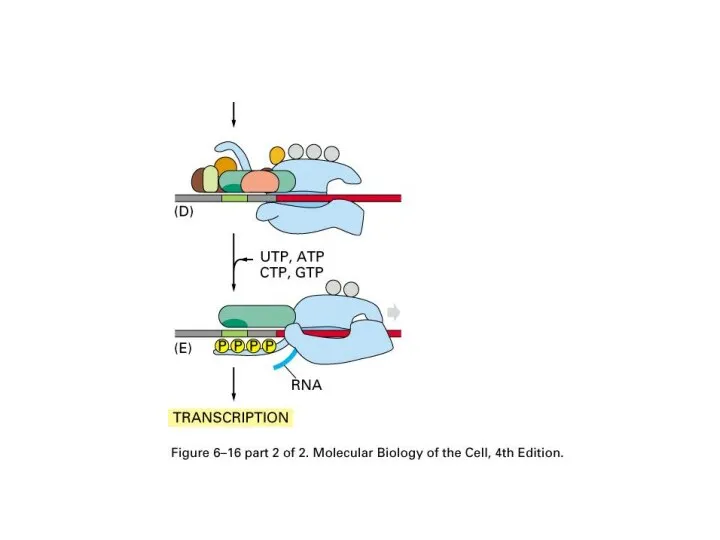
Слайд 32
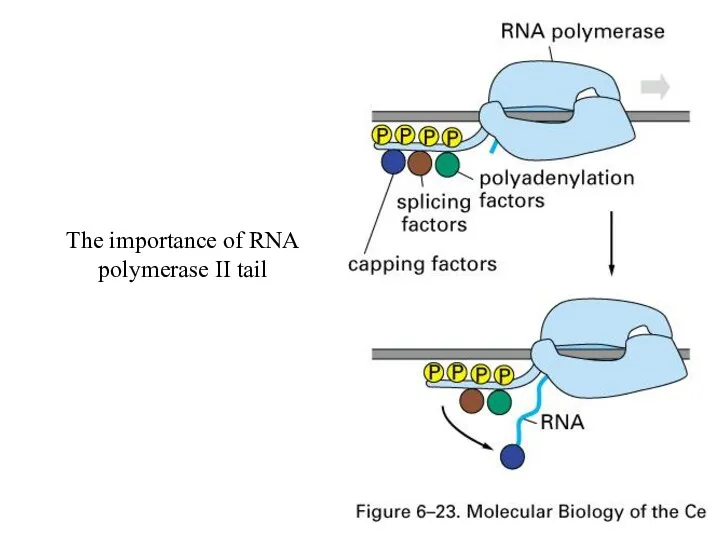
The importance of RNA polymerase II tail
Слайд 33
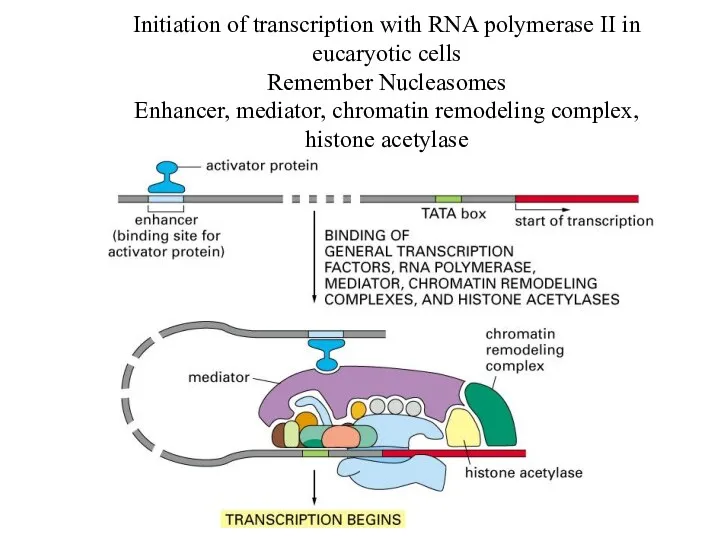
Initiation of transcription with RNA polymerase II in eucaryotic cells
Remember Nucleasomes
Enhancer,
mediator, chromatin remodeling complex, histone acetylase
Слайд 34
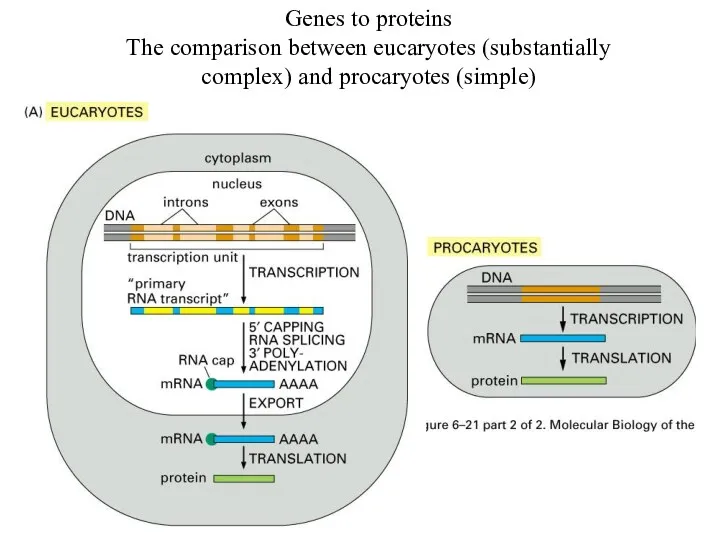
Genes to proteins
The comparison between eucaryotes (substantially complex) and procaryotes (simple)
Слайд 35
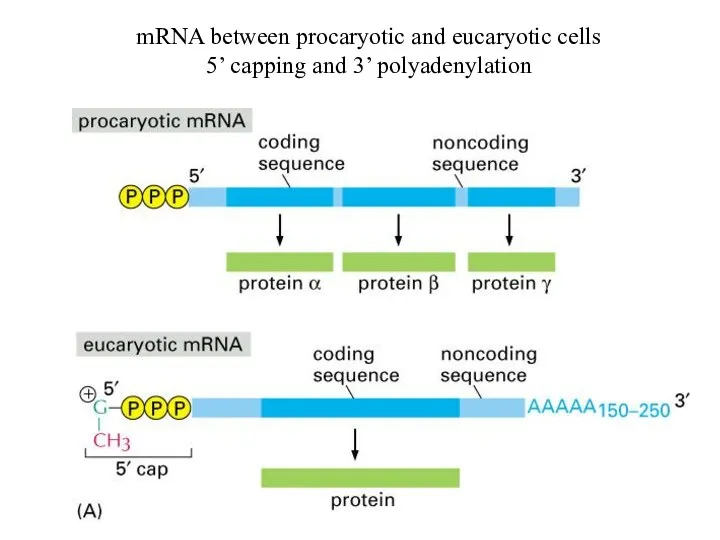
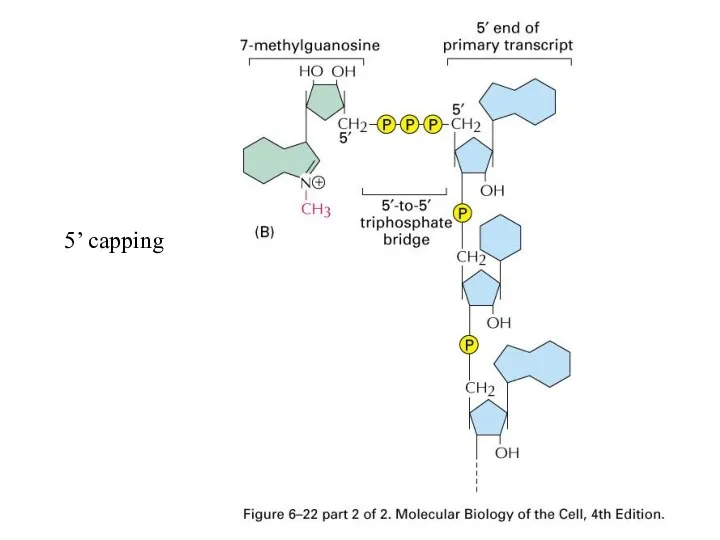
mRNA between procaryotic and eucaryotic cells
5’ capping and 3’ polyadenylation
Слайд 36
Слайд 37
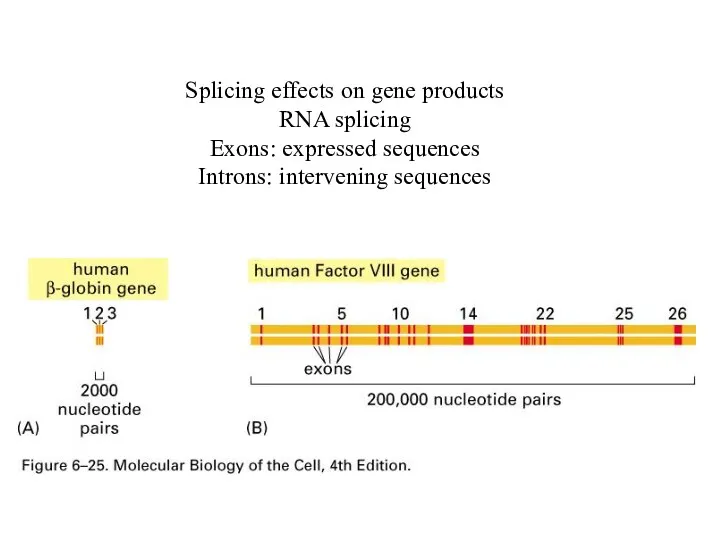
Splicing effects on gene products
RNA splicing
Exons: expressed sequences
Introns: intervening sequences
Слайд 38
Слайд 39
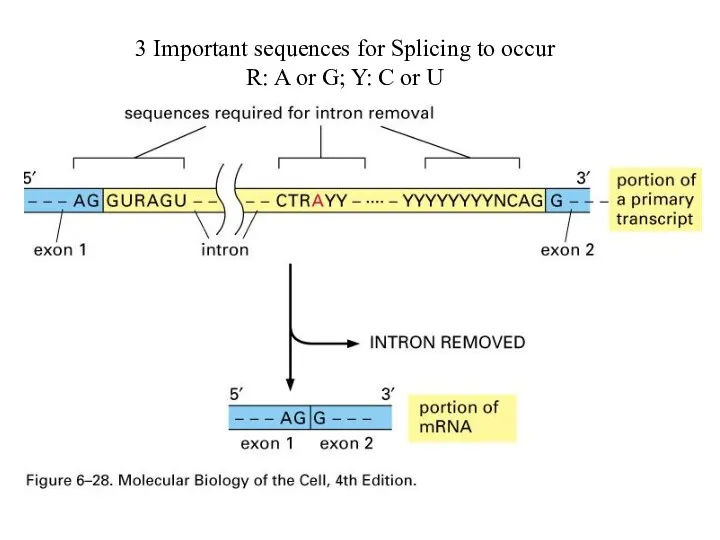
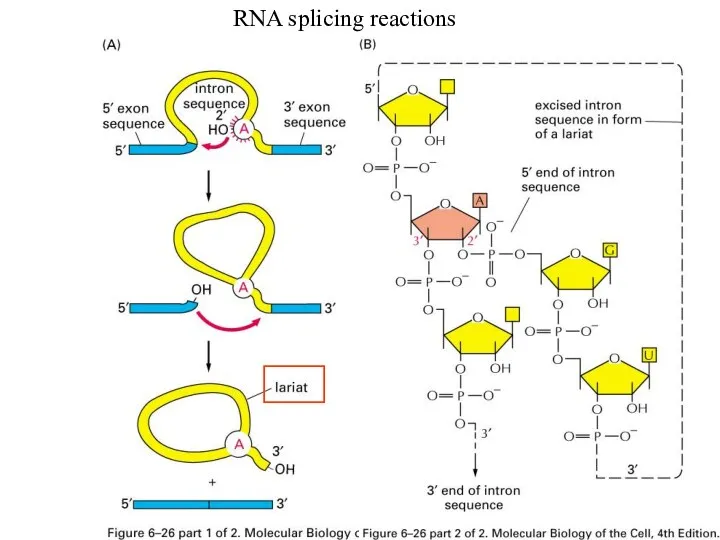
3 Important sequences for Splicing to occur
R: A or G; Y:
C or U
Слайд 40
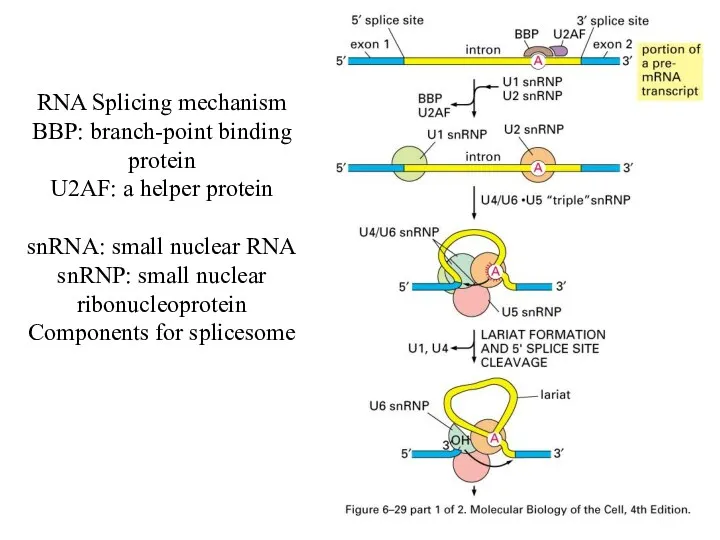
RNA Splicing mechanism
BBP: branch-point binding protein
U2AF: a helper protein
snRNA: small nuclear
RNA
snRNP: small nuclear ribonucleoprotein
Components for splicesome
Слайд 41
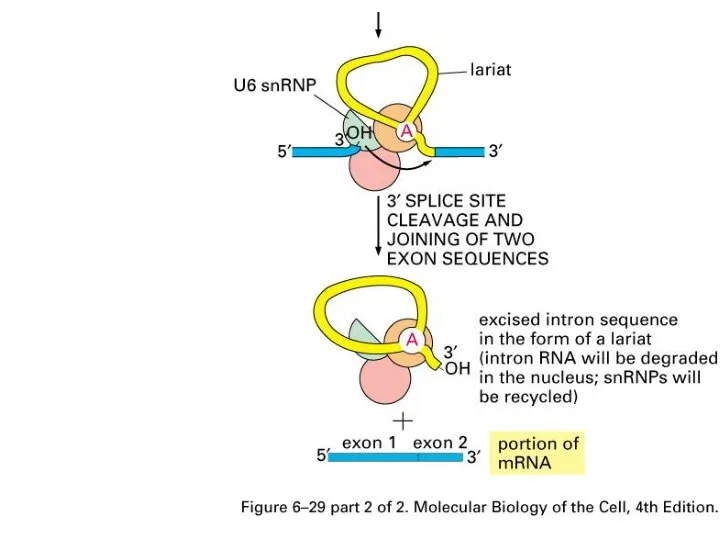
Слайд 42
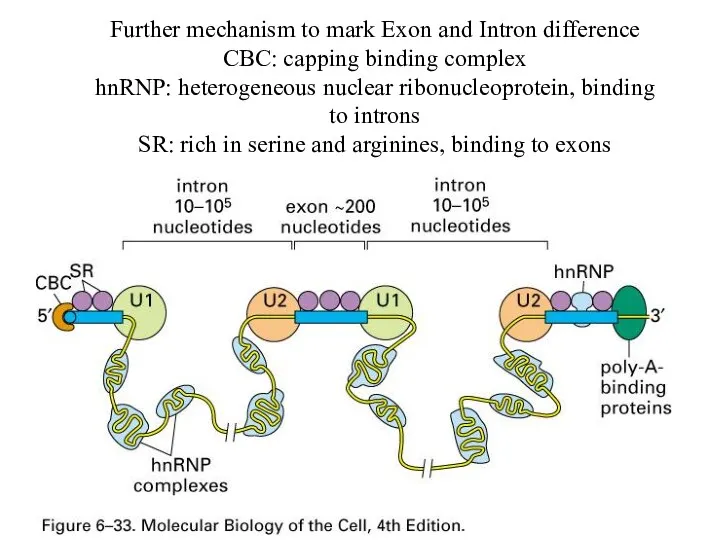
Further mechanism to mark Exon and Intron difference
CBC: capping binding complex
hnRNP:
heterogeneous nuclear ribonucleoprotein, binding to introns
SR: rich in serine and arginines, binding to exons
Слайд 43
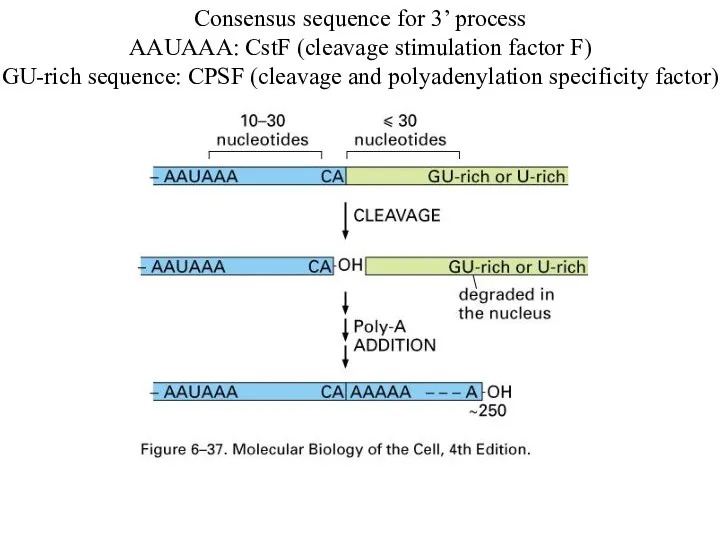
Consensus sequence for 3’ process
AAUAAA: CstF (cleavage stimulation factor F)
GU-rich sequence:
CPSF (cleavage and polyadenylation specificity factor)
Слайд 44
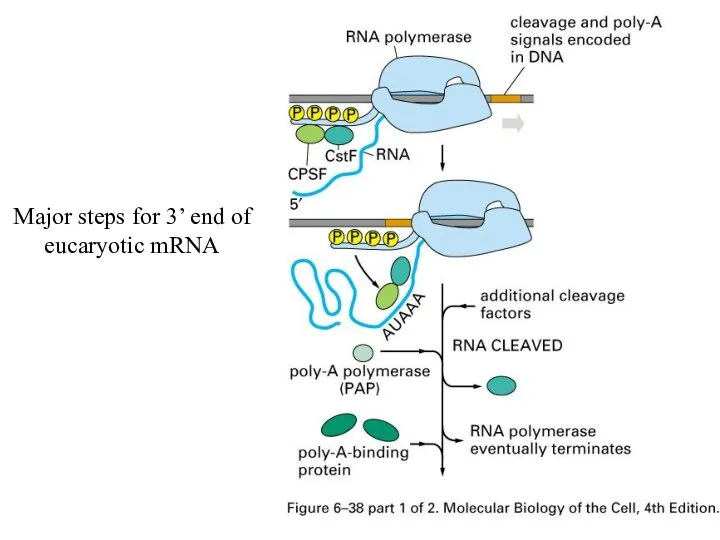
Major steps for 3’ end of eucaryotic mRNA
Слайд 45
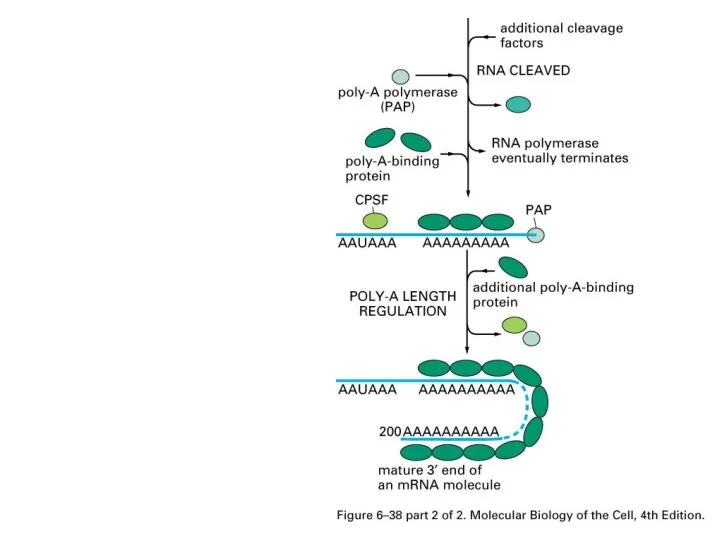
Слайд 46
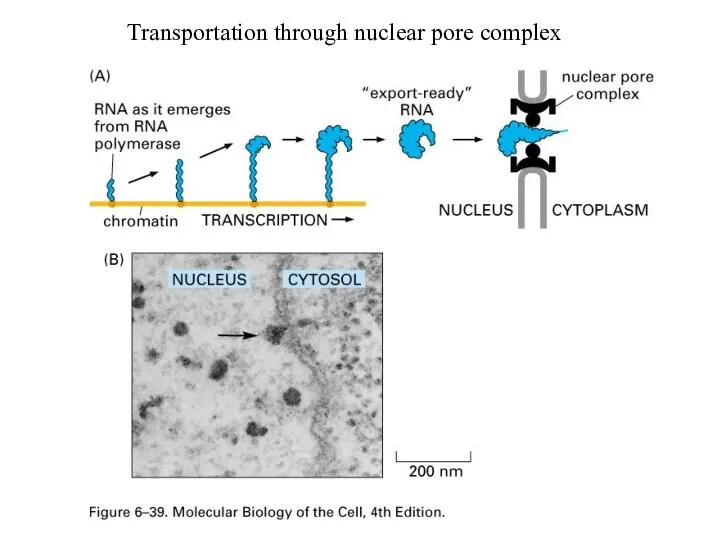
Transportation through nuclear pore complex
Слайд 47
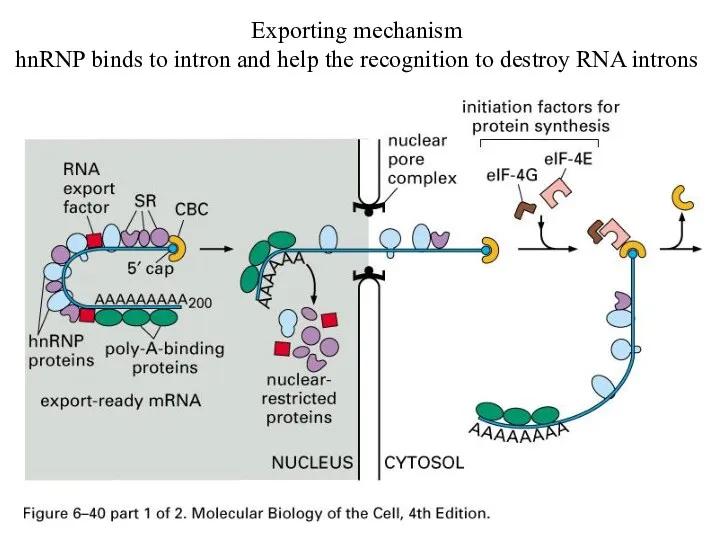
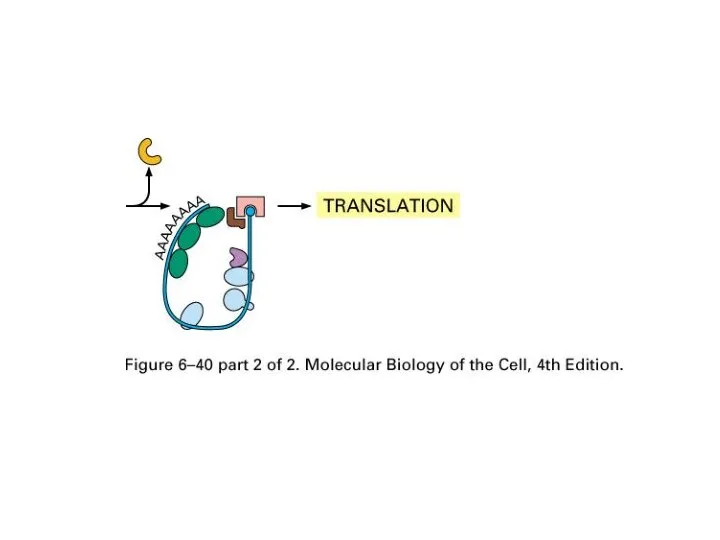
Exporting mechanism
hnRNP binds to intron and help the recognition to destroy
RNA introns
Слайд 48
Слайд 49
Слайд 50
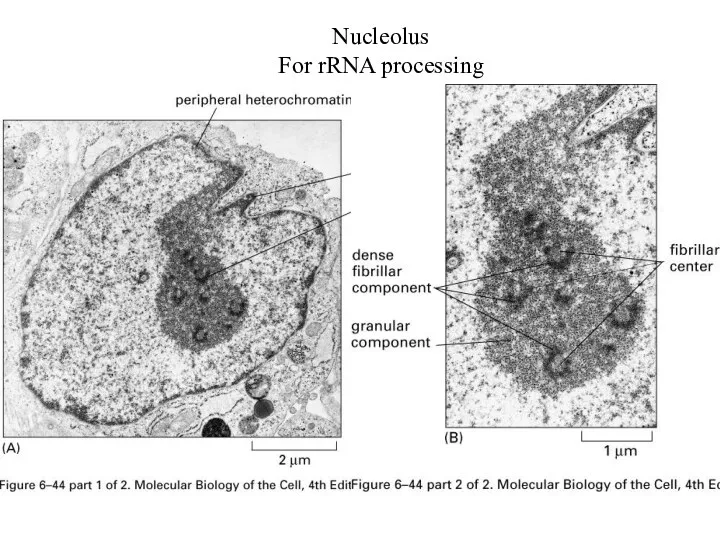
Nucleolus
For rRNA processing
Слайд 51
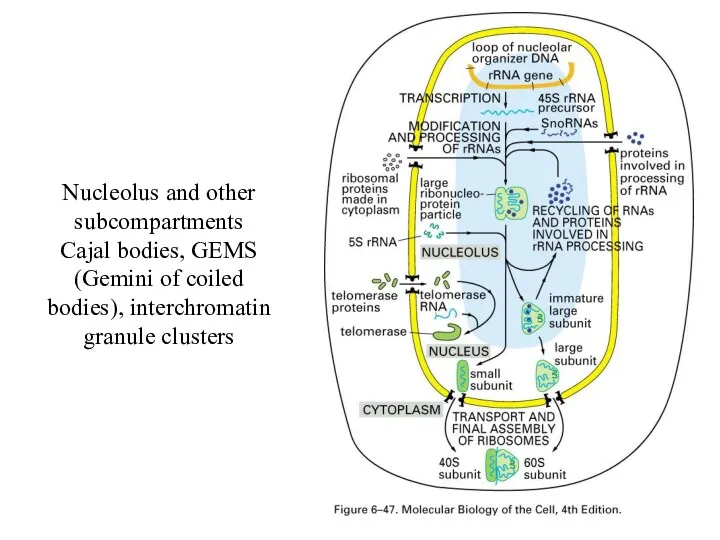
Nucleolus and other subcompartments
Cajal bodies, GEMS (Gemini of coiled bodies), interchromatin
granule clusters
Слайд 52

Summary
Transcription: RNA Polymerase, Promoter, enhancer, transcription factor
5’ capping, splicing, 3’ cleavage
and polyadenylation
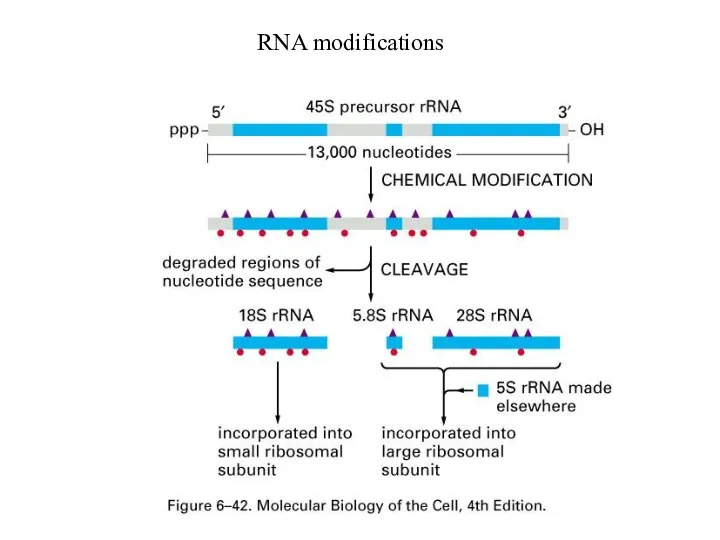
rRNA needs chemical modifications before maturation
Nucleolus with sub-compartments
Слайд 53

From RNA to Protein
Protein synthesis
Protein Folding and regulation
Слайд 54
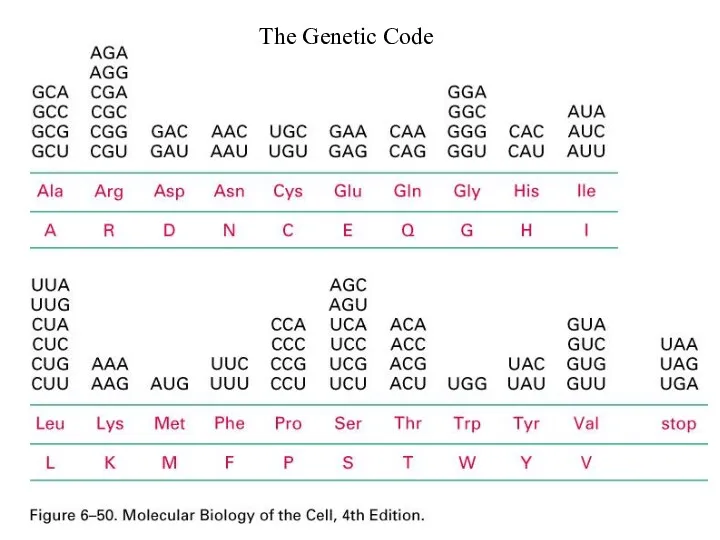
Слайд 55
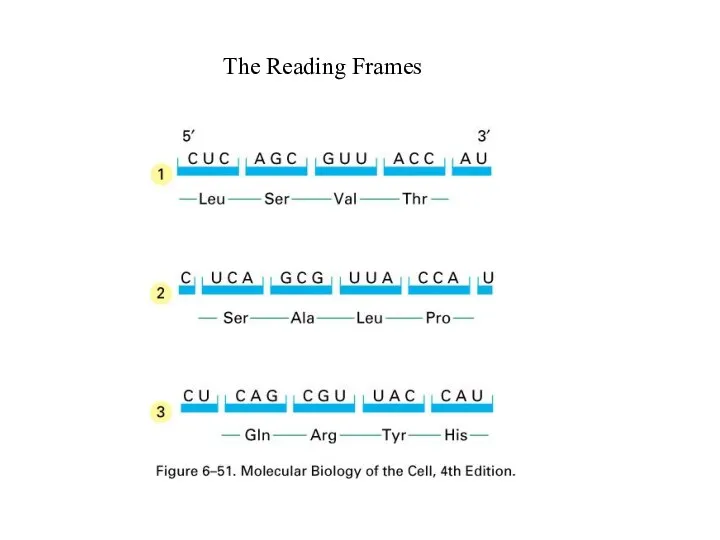
Слайд 56
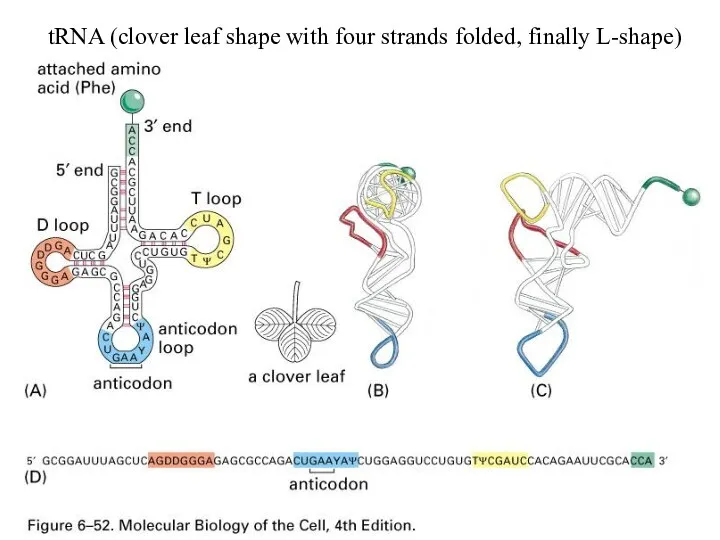
tRNA (clover leaf shape with four strands folded, finally L-shape)
Слайд 57
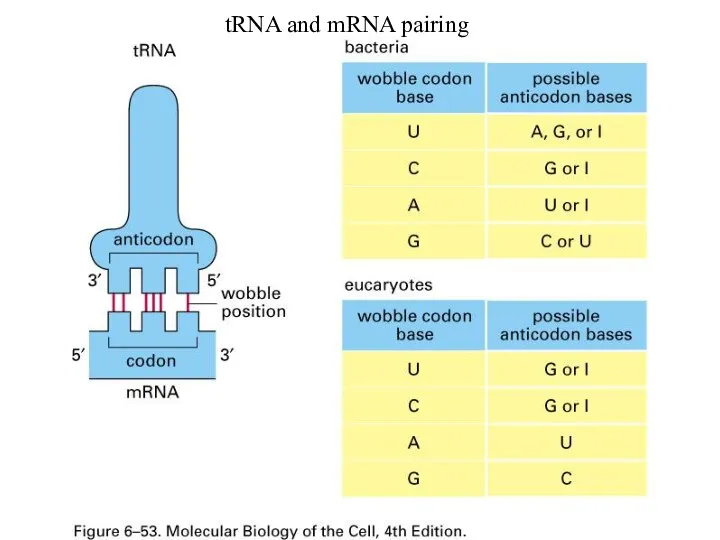
Слайд 58
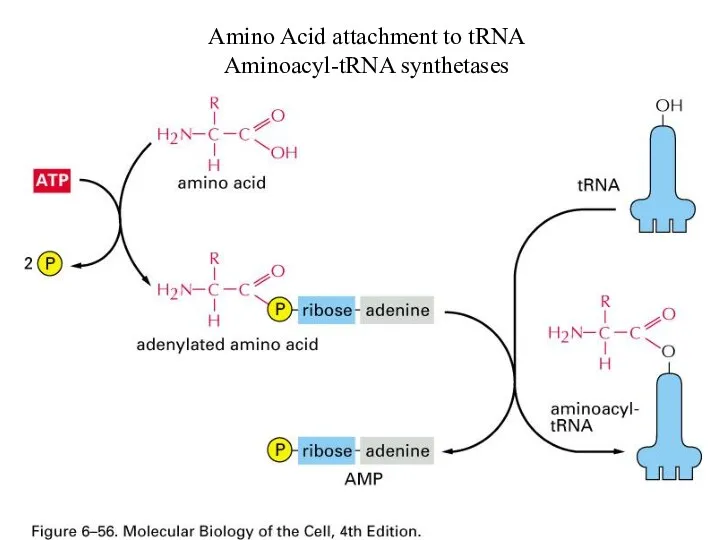
Amino Acid attachment to tRNA
Aminoacyl-tRNA synthetases
Слайд 59
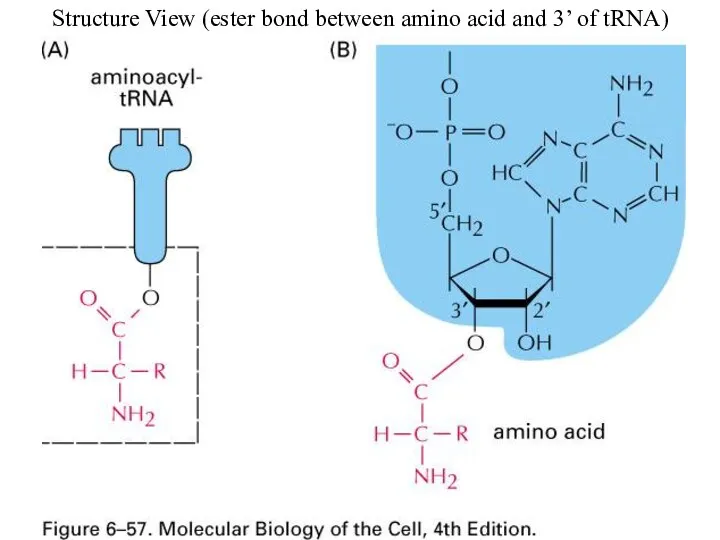
Structure View (ester bond between amino acid and 3’ of tRNA)
Слайд 60
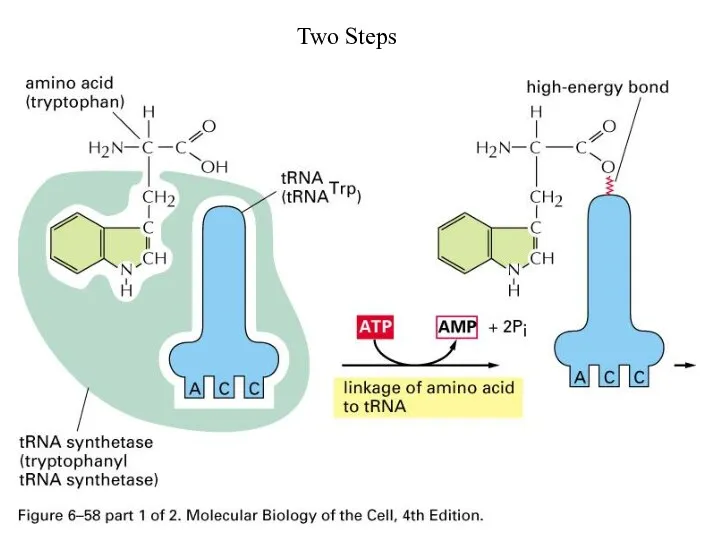
Слайд 61
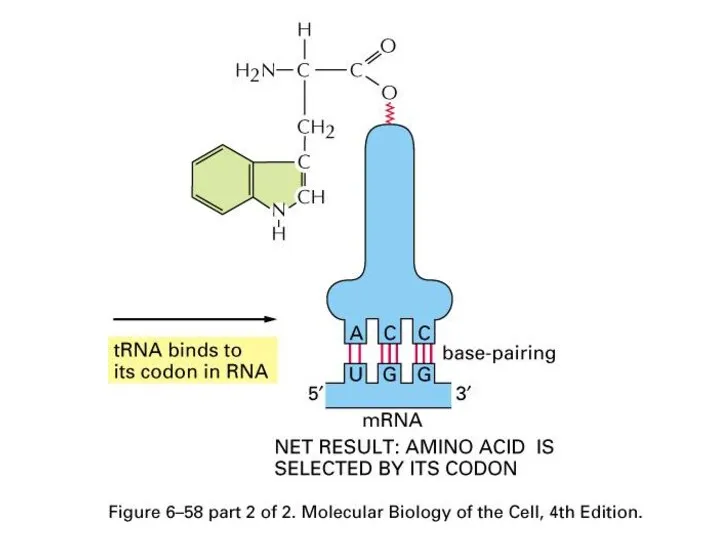
Слайд 62
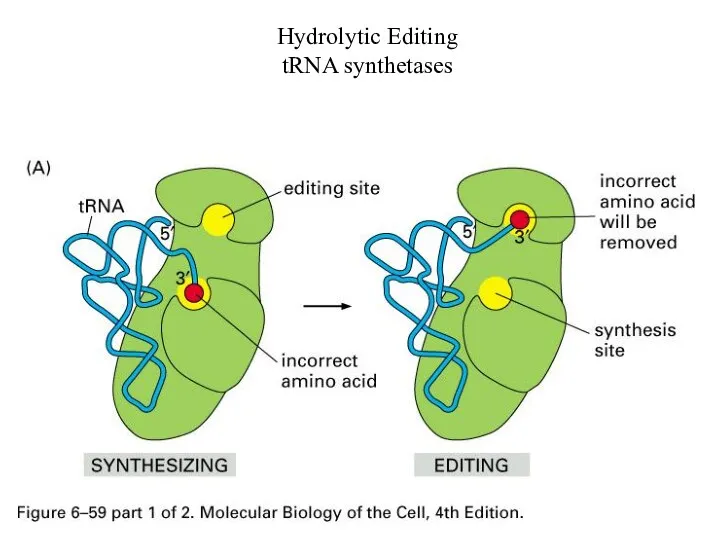
Hydrolytic Editing
tRNA synthetases
Слайд 63
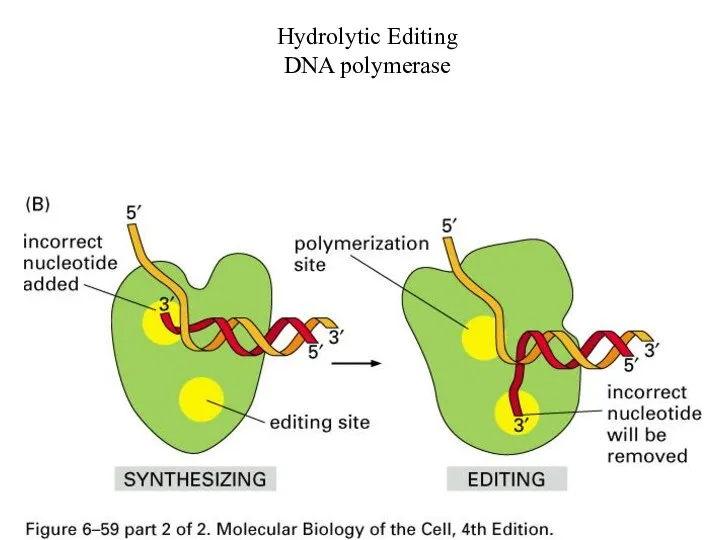
Hydrolytic Editing
DNA polymerase
Слайд 64
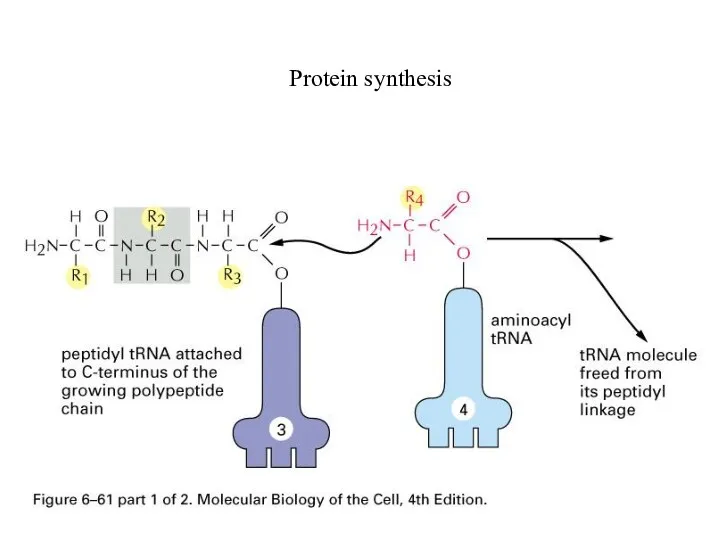
Слайд 65
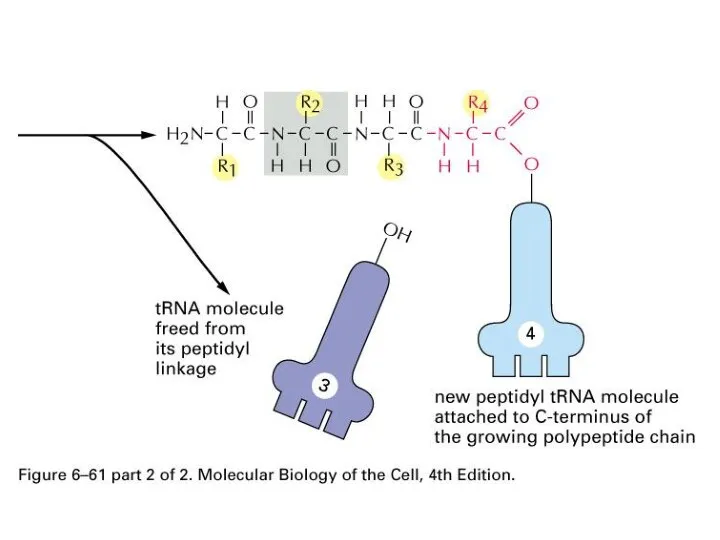
Слайд 66
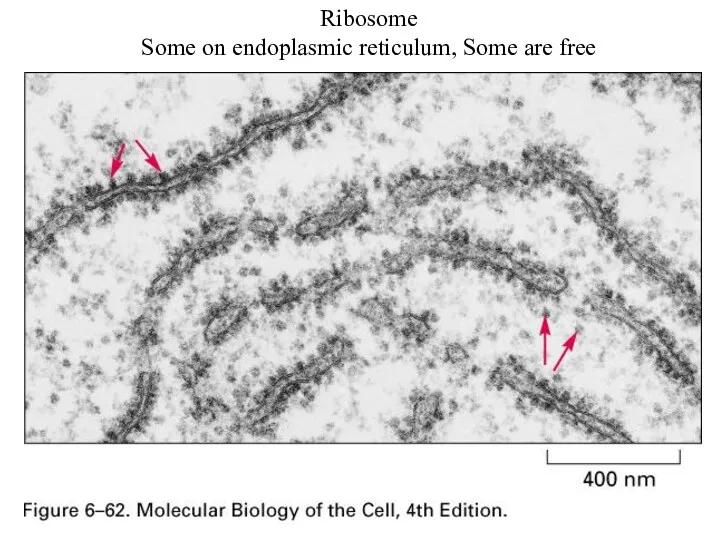
Ribosome
Some on endoplasmic reticulum, Some are free
Слайд 67
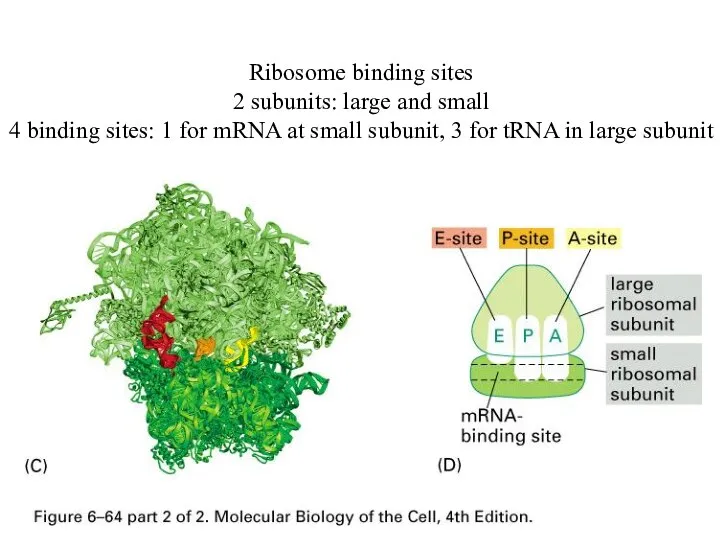
Ribosome binding sites
2 subunits: large and small
4 binding sites: 1 for
mRNA at small subunit, 3 for tRNA in large subunit
Слайд 68
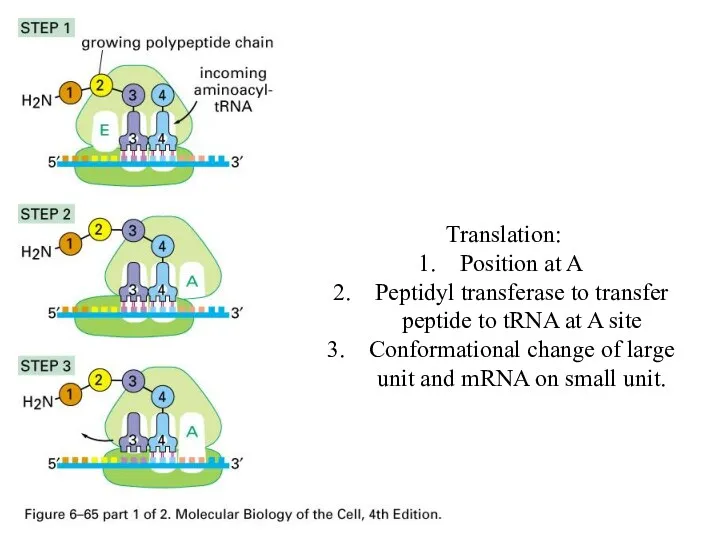
Translation:
Position at A
Peptidyl transferase to transfer peptide to tRNA at A
site
Conformational change of large unit and mRNA on small unit.
Слайд 69
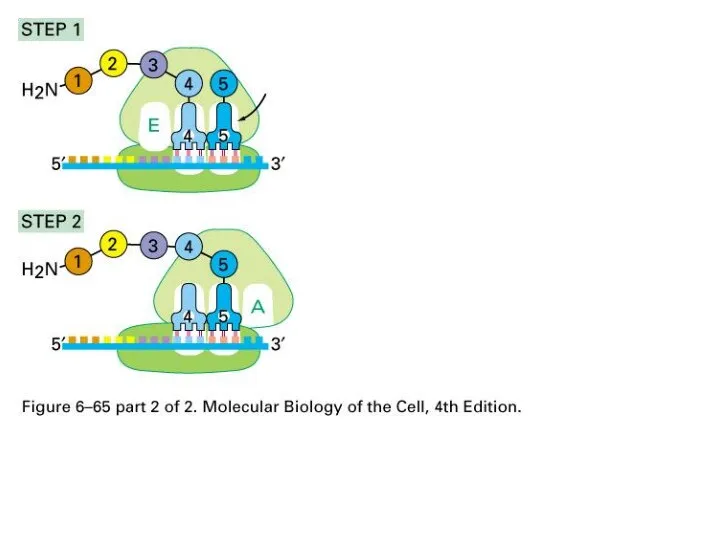
Слайд 70
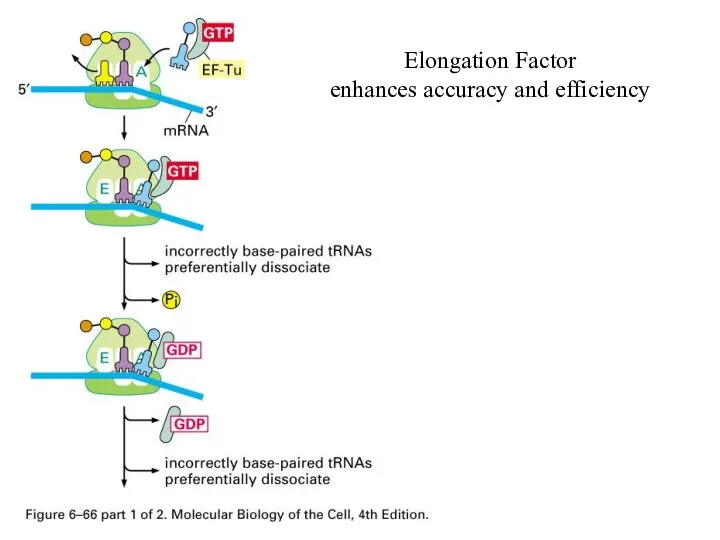
Elongation Factor
enhances accuracy and efficiency
Слайд 71
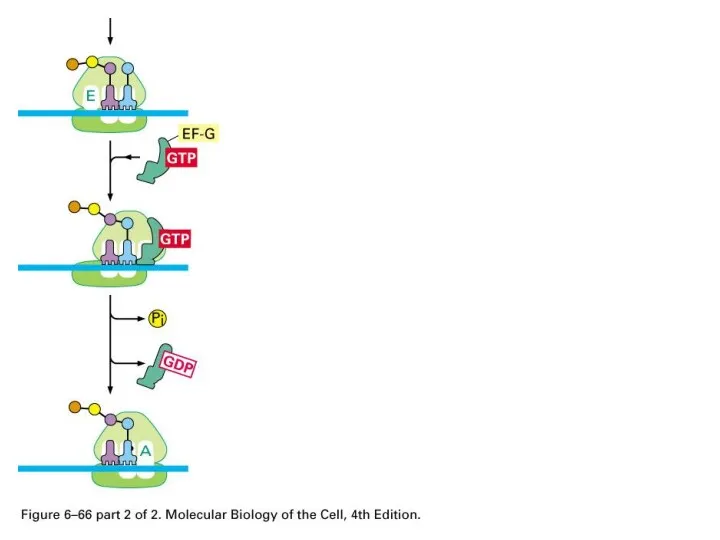
Слайд 72
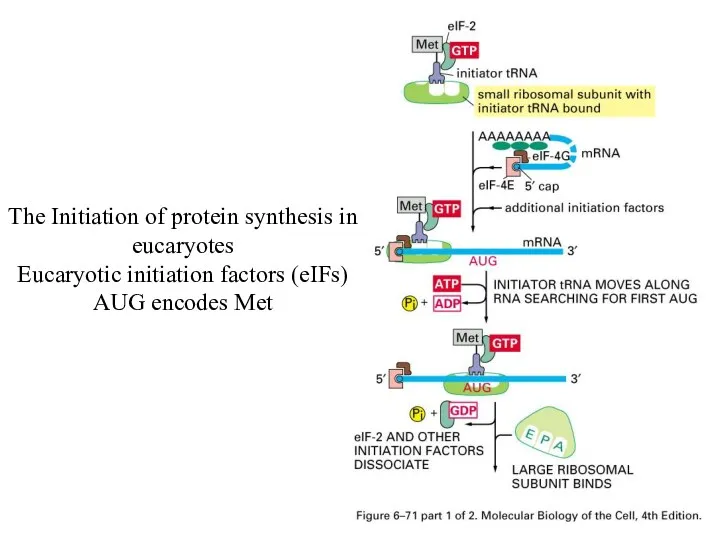
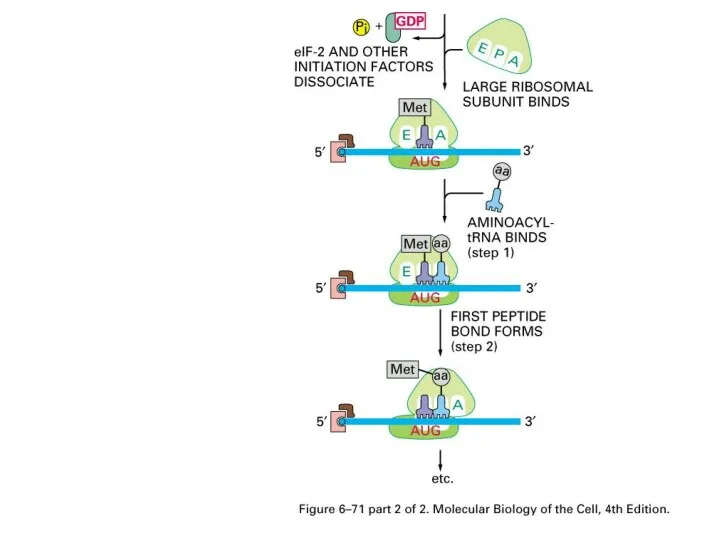
The Initiation of protein synthesis in eucaryotes
Eucaryotic initiation factors (eIFs)
AUG encodes
Met
Слайд 73
Слайд 74
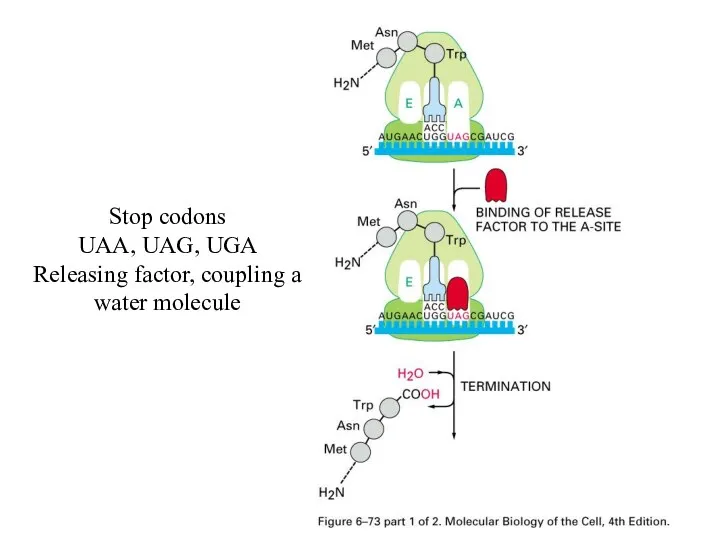
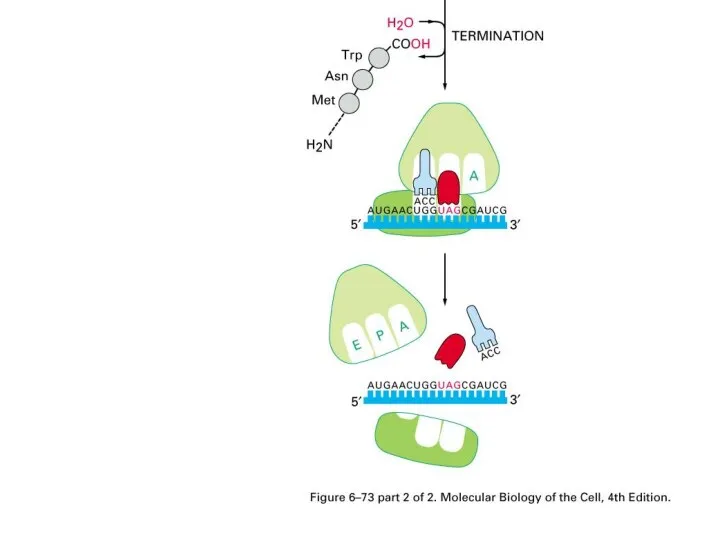
Stop codons
UAA, UAG, UGA
Releasing factor, coupling a water molecule
Слайд 75










































































 Физиология пищеварительной системы и функциональная диагностика
Физиология пищеварительной системы и функциональная диагностика Насекомые
Насекомые Генетика. Наследственность. Изменчивость
Генетика. Наследственность. Изменчивость Пищеварительная система
Пищеварительная система КВН Живая планета
КВН Живая планета Макро- и микроэлементы в питании человека
Макро- и микроэлементы в питании человека Мир динозавров
Мир динозавров Презенетация к уроку Строение цветка. Типы соцветий
Презенетация к уроку Строение цветка. Типы соцветий Мембрани. ранспр
Мембрани. ранспр Урок по биологии на темуГенетика пола
Урок по биологии на темуГенетика пола экскурсия на страусиную ферму
экскурсия на страусиную ферму Покрытосеменные, или Цветковые
Покрытосеменные, или Цветковые Клетка – элементарная единица жизни на земле
Клетка – элементарная единица жизни на земле Декоративные растения в интерьере
Декоративные растения в интерьере Лишайники
Лишайники Аэротенки и их классификация. (Лекция 5.4)
Аэротенки и их классификация. (Лекция 5.4)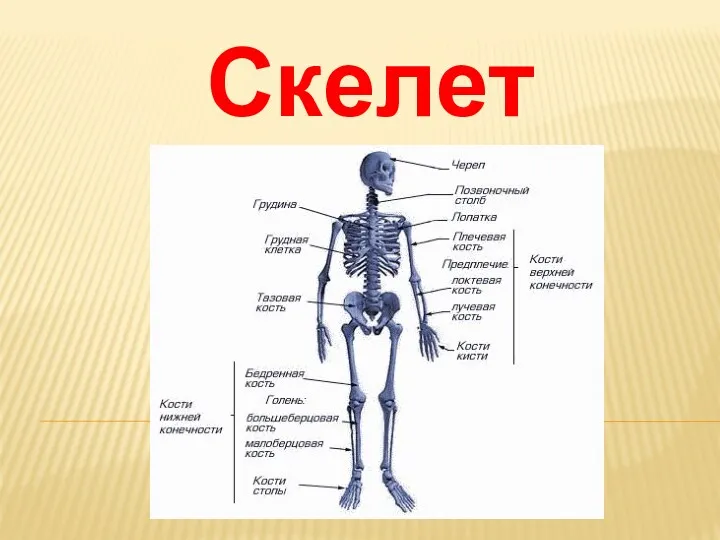 Скелет человека
Скелет человека Вид. Критерии вида
Вид. Критерии вида Вид. Критерии вида. Популяция
Вид. Критерии вида. Популяция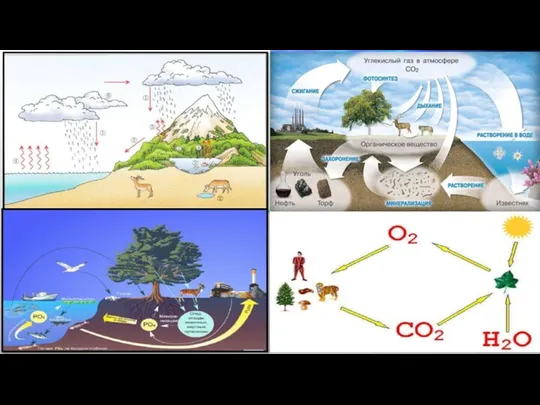 Химический круговорот веществ в природе. 6 класс
Химический круговорот веществ в природе. 6 класс Genetic load of human population
Genetic load of human population Дослідження росту вегетативних органів
Дослідження росту вегетативних органів Дыхание внешнее и внутреннее
Дыхание внешнее и внутреннее Химический состав клетки
Химический состав клетки Бактериологиялық бояулардың түрлерімен танысу
Бактериологиялық бояулардың түрлерімен танысу Чибис - птица 2010 года
Чибис - птица 2010 года Животные Эстонии
Животные Эстонии Як тварини використовують знаряддя праці
Як тварини використовують знаряддя праці